| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
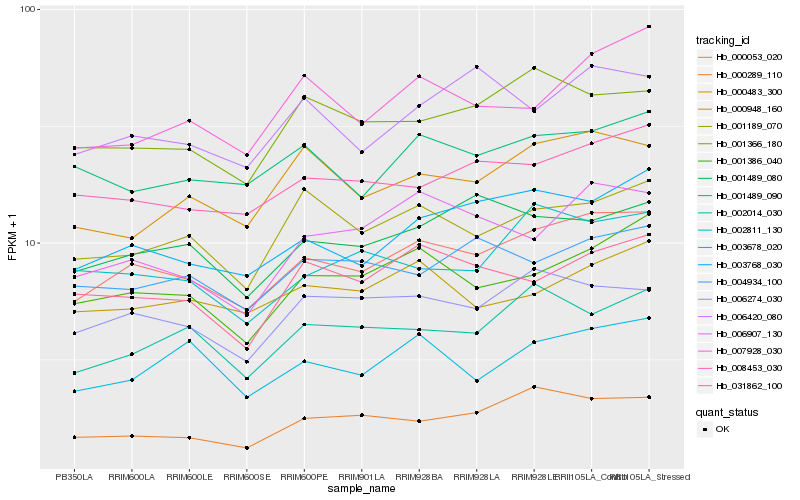
Hb_000053_020 |
0.0 |
- |
- |
F-box and wd40 domain protein, putative [Ricinus communis] |
| 2 |
Hb_003768_030 |
0.06578208 |
- |
- |
PREDICTED: ribosome biogenesis protein WDR12 homolog isoform X1 [Jatropha curcas] |
| 3 |
Hb_008453_030 |
0.0713730513 |
- |
- |
Thioredoxin superfamily protein [Theobroma cacao] |
| 4 |
Hb_001386_040 |
0.0726860192 |
transcription factor |
TF Family: C2H2 |
PREDICTED: zinc finger protein 511 [Jatropha curcas] |
| 5 |
Hb_006274_030 |
0.0743581029 |
- |
- |
F-box and wd40 domain protein, putative [Ricinus communis] |
| 6 |
Hb_001189_070 |
0.079401103 |
- |
- |
PREDICTED: ribosome production factor 1 [Jatropha curcas] |
| 7 |
Hb_001489_090 |
0.0798070964 |
- |
- |
PREDICTED: uncharacterized protein LOC105649775 [Jatropha curcas] |
| 8 |
Hb_000483_300 |
0.0818531178 |
- |
- |
PREDICTED: ATP-dependent Clp protease proteolytic subunit-related protein 4, chloroplastic [Populus euphratica] |
| 9 |
Hb_001366_180 |
0.0822671756 |
- |
- |
PREDICTED: translocase of chloroplast 33, chloroplastic-like [Jatropha curcas] |
| 10 |
Hb_031862_100 |
0.0826310656 |
- |
- |
pentatricopeptide repeat-containing protein, putative [Ricinus communis] |
| 11 |
Hb_002014_030 |
0.0834888694 |
- |
- |
hypothetical protein JCGZ_07251 [Jatropha curcas] |
| 12 |
Hb_001489_080 |
0.084252981 |
- |
- |
Protein kinase APK1B, chloroplast precursor, putative [Ricinus communis] |
| 13 |
Hb_000948_160 |
0.0845455539 |
- |
- |
PREDICTED: 1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino] imidazole-4-carboxamide isomerase, chloroplastic [Jatropha curcas] |
| 14 |
Hb_006907_130 |
0.0851771795 |
- |
- |
unnamed protein product [Coffea canephora] |
| 15 |
Hb_003678_020 |
0.0855420693 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 16 |
Hb_006420_080 |
0.0867120062 |
rubber biosynthesis |
Gene Name: Pyruvate dehydrogenase |
PREDICTED: pyruvate dehydrogenase E1 component subunit alpha-1, mitochondrial isoform X1 [Jatropha curcas] |
| 17 |
Hb_007928_030 |
0.086767515 |
- |
- |
PREDICTED: dual specificity phosphatase Cdc25 [Jatropha curcas] |
| 18 |
Hb_000289_110 |
0.0868045986 |
- |
- |
PREDICTED: cysteine synthase-like [Musa acuminata subsp. malaccensis] |
| 19 |
Hb_004934_100 |
0.0868400169 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 20 |
Hb_002811_130 |
0.0869813842 |
- |
- |
Zeta-carotene desaturase, chloroplast precursor, putative [Ricinus communis] |