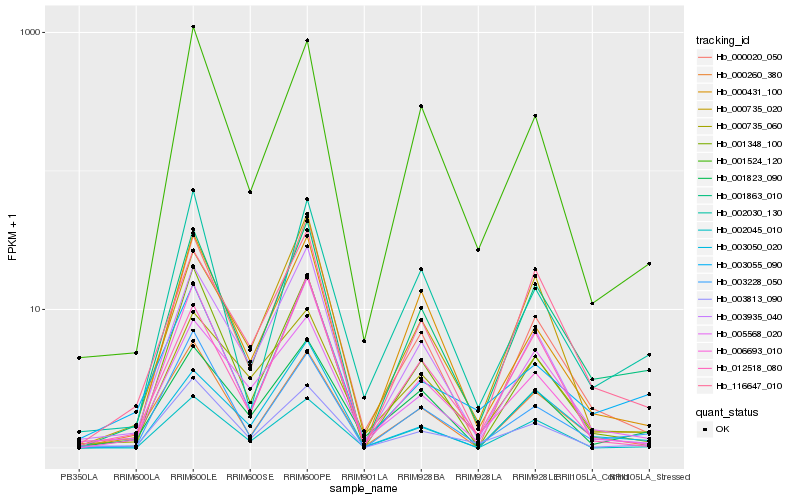
| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_001863_010 |
0.0 |
- |
- |
PREDICTED: classical arabinogalactan protein 4-like [Jatropha curcas] |
| 2 |
Hb_116647_010 |
0.1154928154 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 3 |
Hb_001524_120 |
0.134023063 |
- |
- |
myo-inositol-1 phosphate synthase [Hevea brasiliensis] |
| 4 |
Hb_000260_380 |
0.1372363872 |
- |
- |
PREDICTED: cell division control protein 2 homolog C [Jatropha curcas] |
| 5 |
Hb_003228_050 |
0.1380037091 |
- |
- |
chromomethylase [Hevea brasiliensis] |
| 6 |
Hb_000431_100 |
0.1398492581 |
- |
- |
ribonucleoside-diphosphate reductase small chain, putative [Ricinus communis] |
| 7 |
Hb_003050_020 |
0.1407414194 |
- |
- |
glycosyl hydrolase family 17 family protein [Populus trichocarpa] |
| 8 |
Hb_001823_090 |
0.1409240354 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 9 |
Hb_002030_130 |
0.1438812356 |
- |
- |
hypothetical protein CARUB_v10021660mg, partial [Capsella rubella] |
| 10 |
Hb_003813_090 |
0.1445427804 |
- |
- |
PREDICTED: uncharacterized protein LOC105638743 [Jatropha curcas] |
| 11 |
Hb_000020_050 |
0.1454612402 |
- |
- |
PREDICTED: expansin-A6 [Jatropha curcas] |
| 12 |
Hb_000735_060 |
0.1457092137 |
- |
- |
PREDICTED: U-box domain-containing protein 9 [Jatropha curcas] |
| 13 |
Hb_001348_100 |
0.1469624608 |
- |
- |
PREDICTED: uncharacterized protein LOC105638934 [Jatropha curcas] |
| 14 |
Hb_003935_040 |
0.1473481345 |
transcription factor |
TF Family: MIKC |
PREDICTED: MADS-box protein SVP [Jatropha curcas] |
| 15 |
Hb_003055_090 |
0.1475984001 |
- |
- |
PREDICTED: uncharacterized protein LOC105649490 [Jatropha curcas] |
| 16 |
Hb_005568_020 |
0.1496795104 |
transcription factor |
TF Family: ERF |
PREDICTED: ethylene-responsive transcription factor RAP2-7 isoform X1 [Jatropha curcas] |
| 17 |
Hb_012518_080 |
0.1500054782 |
- |
- |
PREDICTED: glucan endo-1,3-beta-glucosidase 11-like [Jatropha curcas] |
| 18 |
Hb_000735_020 |
0.1521505329 |
- |
- |
Auxin-induced in root cultures protein 12 precursor, putative [Ricinus communis] |
| 19 |
Hb_006693_010 |
0.1538352429 |
- |
- |
PREDICTED: uncharacterized protein LOC105628875 [Jatropha curcas] |
| 20 |
Hb_002045_010 |
0.1541798198 |
- |
- |
PREDICTED: F-box/LRR-repeat protein 17 [Jatropha curcas] |