| Rank |
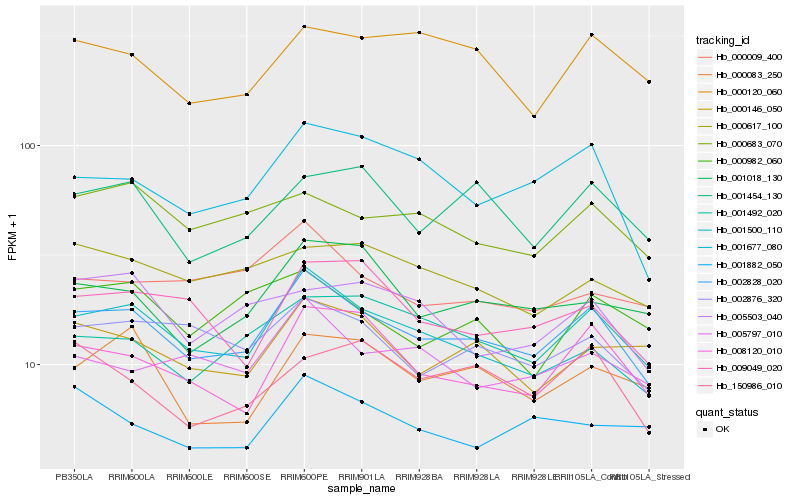
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_002828_020 |
0.0 |
- |
- |
protein binding protein, putative [Ricinus communis] |
| 2 |
Hb_001500_110 |
0.0691859182 |
- |
- |
serine-threonine protein kinase, plant-type, putative [Ricinus communis] |
| 3 |
Hb_008120_010 |
0.0809465421 |
- |
- |
PREDICTED: polyadenylate-binding protein-interacting protein 9-like [Populus euphratica] |
| 4 |
Hb_002876_320 |
0.0840893261 |
- |
- |
PREDICTED: uncharacterized protein LOC105632052 [Jatropha curcas] |
| 5 |
Hb_001677_080 |
0.0868331257 |
- |
- |
mannose-1-phosphate guanyltransferase, putative [Ricinus communis] |
| 6 |
Hb_001492_020 |
0.0919131076 |
- |
- |
PREDICTED: diacylglycerol kinase 3-like [Jatropha curcas] |
| 7 |
Hb_000683_070 |
0.0924274356 |
- |
- |
d-3-phosphoglycerate dehydrogenase, putative [Ricinus communis] |
| 8 |
Hb_005797_010 |
0.0937291327 |
- |
- |
PREDICTED: lysophospholipid acyltransferase 1-like [Jatropha curcas] |
| 9 |
Hb_001454_130 |
0.0938042522 |
- |
- |
Xyloglucan 6-xylosyltransferase, putative [Ricinus communis] |
| 10 |
Hb_001018_130 |
0.0941802974 |
- |
- |
PREDICTED: uncharacterized protein LOC105638260 [Jatropha curcas] |
| 11 |
Hb_009049_020 |
0.0957786268 |
- |
- |
PREDICTED: 2-dehydro-3-deoxyphosphooctonate aldolase 1-like [Jatropha curcas] |
| 12 |
Hb_000146_050 |
0.0959054481 |
- |
- |
PREDICTED: protein unc-50 homolog isoform X2 [Gossypium raimondii] |
| 13 |
Hb_000120_060 |
0.0974120791 |
- |
- |
PREDICTED: peptidyl-prolyl cis-trans isomerase CYP20-1 [Populus euphratica] |
| 14 |
Hb_000009_400 |
0.0989414012 |
- |
- |
PREDICTED: BTB/POZ domain-containing protein At3g50780 [Jatropha curcas] |
| 15 |
Hb_000617_100 |
0.0998106128 |
- |
- |
PREDICTED: uncharacterized protein LOC105647493 [Jatropha curcas] |
| 16 |
Hb_000083_250 |
0.0998774235 |
- |
- |
ring finger protein, putative [Ricinus communis] |
| 17 |
Hb_001882_050 |
0.1022685529 |
- |
- |
hydroxysteroid dehydrogenase, putative [Ricinus communis] |
| 18 |
Hb_150986_010 |
0.1024230196 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 19 |
Hb_000982_060 |
0.1029089597 |
- |
- |
PREDICTED: probable protein S-acyltransferase 14 [Jatropha curcas] |
| 20 |
Hb_005503_040 |
0.1041495241 |
- |
- |
PREDICTED: AMP deaminase [Jatropha curcas] |