| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
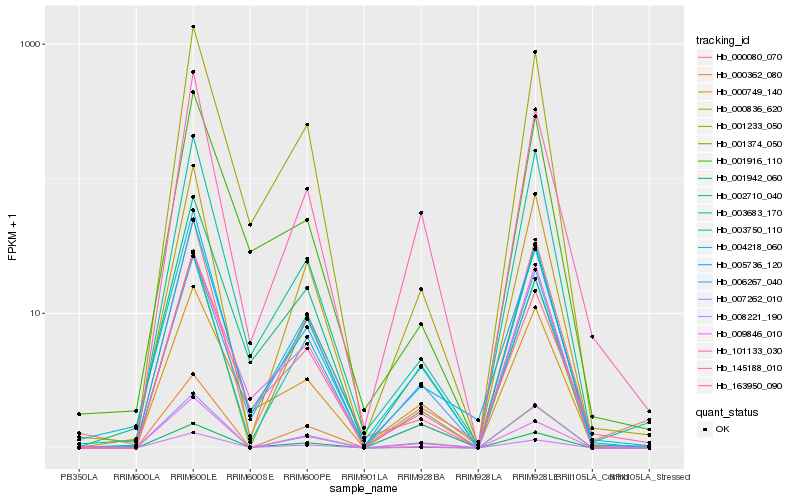
Hb_004218_060 |
0.0 |
- |
- |
annexin [Manihot esculenta] |
| 2 |
Hb_005736_120 |
0.0822130347 |
- |
- |
PREDICTED: carboxyvinyl-carboxyphosphonate phosphorylmutase, chloroplastic [Gossypium raimondii] |
| 3 |
Hb_145188_010 |
0.086269467 |
- |
- |
PREDICTED: endochitinase PR4-like [Jatropha curcas] |
| 4 |
Hb_002710_040 |
0.0936927792 |
- |
- |
PREDICTED: cyclin-D3-3-like [Jatropha curcas] |
| 5 |
Hb_001942_060 |
0.0941978309 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 6 |
Hb_007262_010 |
0.0990363508 |
- |
- |
PREDICTED: CCR4-NOT transcription complex subunit 10 isoform X2 [Jatropha curcas] |
| 7 |
Hb_163950_090 |
0.0994155008 |
- |
- |
PREDICTED: traB domain-containing protein isoform X1 [Jatropha curcas] |
| 8 |
Hb_003750_110 |
0.105934515 |
- |
- |
PREDICTED: aldose 1-epimerase-like [Jatropha curcas] |
| 9 |
Hb_000362_080 |
0.1066091954 |
- |
- |
hypothetical protein EUTSA_v10027329mg, partial [Eutrema salsugineum] |
| 10 |
Hb_000749_140 |
0.1068776904 |
- |
- |
PREDICTED: 2-succinylbenzoate--CoA ligase, chloroplastic/peroxisomal [Jatropha curcas] |
| 11 |
Hb_000836_620 |
0.1117819517 |
- |
- |
PREDICTED: protochlorophyllide reductase, chloroplastic [Jatropha curcas] |
| 12 |
Hb_006267_040 |
0.1133155 |
- |
- |
PREDICTED: ABC transporter G family member 32 isoform X1 [Jatropha curcas] |
| 13 |
Hb_001233_050 |
0.1134098968 |
- |
- |
Magnesium-protoporphyrin IX monomethyl ester [oxidative] cyclase, chloroplast precursor, putative [Ricinus communis] |
| 14 |
Hb_000080_070 |
0.1165795045 |
- |
- |
PREDICTED: peptidyl-prolyl cis-trans isomerase FKBP16-3, chloroplastic [Jatropha curcas] |
| 15 |
Hb_003683_170 |
0.1179300504 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 16 |
Hb_009846_010 |
0.1195420819 |
- |
- |
PREDICTED: uncharacterized protein LOC105643801 [Jatropha curcas] |
| 17 |
Hb_101133_030 |
0.1197492145 |
- |
- |
DUF26 domain-containing protein 1 precursor, putative [Ricinus communis] |
| 18 |
Hb_008221_190 |
0.1220854766 |
- |
- |
PREDICTED: probable LRR receptor-like serine/threonine-protein kinase At1g53440 [Jatropha curcas] |
| 19 |
Hb_001916_110 |
0.1243493757 |
- |
- |
PREDICTED: uncharacterized protein LOC105641510 [Jatropha curcas] |
| 20 |
Hb_001374_050 |
0.1260259917 |
- |
- |
PREDICTED: subtilisin-like protease SBT5.3 [Jatropha curcas] |