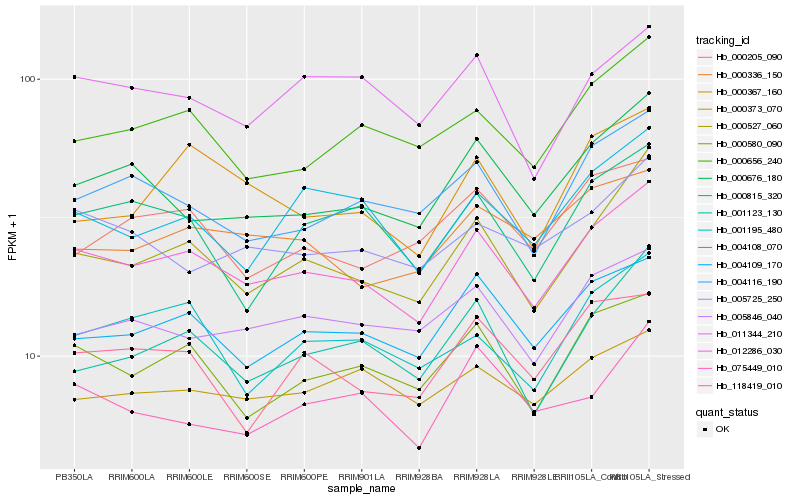
| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_012286_030 |
0.0 |
transcription factor |
TF Family: MED7 |
PREDICTED: mediator of RNA polymerase II transcription subunit 7a [Solanum lycopersicum] |
| 2 |
Hb_000527_060 |
0.0504464924 |
- |
- |
Protein C20orf11, putative [Ricinus communis] |
| 3 |
Hb_000580_090 |
0.0581251008 |
- |
- |
PREDICTED: tRNA 2'-phosphotransferase 1 isoform X1 [Jatropha curcas] |
| 4 |
Hb_004109_170 |
0.0588615057 |
- |
- |
PREDICTED: probable ribokinase [Jatropha curcas] |
| 5 |
Hb_000676_180 |
0.0685603913 |
- |
- |
Ubiquitin-conjugating enzyme 16 [Theobroma cacao] |
| 6 |
Hb_004116_190 |
0.0696046686 |
transcription factor |
TF Family: Alfin-like |
PREDICTED: PHD finger protein ALFIN-LIKE 2 [Jatropha curcas] |
| 7 |
Hb_005846_040 |
0.0704601983 |
- |
- |
PREDICTED: ubiquitin carboxyl-terminal hydrolase 4 [Populus euphratica] |
| 8 |
Hb_000656_240 |
0.0709025655 |
- |
- |
proteasome subunit alpha type, putative [Ricinus communis] |
| 9 |
Hb_004108_070 |
0.0709614033 |
- |
- |
PREDICTED: BTB/POZ and MATH domain-containing protein 2-like [Jatropha curcas] |
| 10 |
Hb_011344_210 |
0.0728467584 |
- |
- |
PREDICTED: probable calcium-binding protein CML13 [Jatropha curcas] |
| 11 |
Hb_001123_130 |
0.0728803546 |
transcription factor |
TF Family: C3H |
nucleic acid binding protein, putative [Ricinus communis] |
| 12 |
Hb_005725_250 |
0.0730340348 |
- |
- |
PREDICTED: uncharacterized protein LOC105630805 isoform X1 [Jatropha curcas] |
| 13 |
Hb_001195_480 |
0.0731739659 |
- |
- |
cop9 complex subunit, putative [Ricinus communis] |
| 14 |
Hb_000336_150 |
0.0736605017 |
- |
- |
ribulose-5-phosphate-3-epimerase, putative [Ricinus communis] |
| 15 |
Hb_118419_010 |
0.0759453825 |
transcription factor |
TF Family: FAR1 |
PREDICTED: protein FAR1-RELATED SEQUENCE 5 [Jatropha curcas] |
| 16 |
Hb_000815_320 |
0.0767112373 |
- |
- |
PREDICTED: vesicle transport v-SNARE 12-like isoform X2 [Jatropha curcas] |
| 17 |
Hb_000373_070 |
0.0774299278 |
- |
- |
PREDICTED: serine/threonine-protein kinase HT1-like isoform X2 [Jatropha curcas] |
| 18 |
Hb_000367_160 |
0.0776259506 |
- |
- |
PREDICTED: SNAP25 homologous protein SNAP33 [Populus euphratica] |
| 19 |
Hb_075449_010 |
0.0798859521 |
- |
- |
PREDICTED: cysteine synthase 2-like [Jatropha curcas] |
| 20 |
Hb_000205_090 |
0.0802379469 |
- |
- |
hypothetical protein POPTR_0019s03570g [Populus trichocarpa] |